On Wednesday, the Nobel Committee announced that it had awarded the Nobel Prize in chemistry to researchers who pioneered major breakthroughs in computational chemistry. These include two researchers at Google’s DeepMind in acknowledgment of their role in developing AI software that could take a raw protein sequence and use it to predict the three-dimensional structure the protein would adopt in cells. Separately, the University of Washington’s David Baker was honored for developing software that could design entirely new proteins with specific structures.
The award makes for a bit of a theme for this year, as yesterday’s Physics prize honored AI developments. In that case, the connection to physics seemed a bit tenuous, but here, there should be little question that the developments solved major problems in biochemistry.
Understanding protein structure
DeepMind, represented by Demis Hassabis and John Jumper, had developed AIs that managed to master games as diverse as chess and StarCraft. But it was always working on more significant problems in parallel, and in 2020, it surprised many people by announcing that it had tackled one of the biggest computational challenges in existence: the prediction of protein structures.
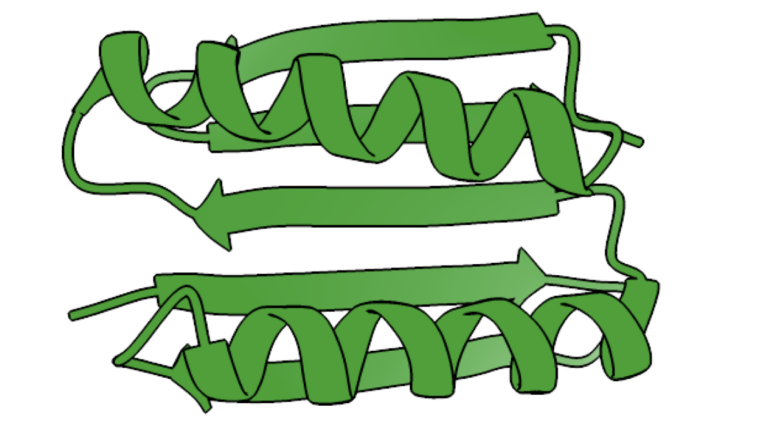
Chemically, proteins are a linear string of amino acids linked together, with living creatures typically having the choice of 20 different amino acids for each position along the string. Most of those 20 have distinctive chemical properties: some are acidic, others basic; some may be negatively charged, others positively charged, and still others neutral, etc. These properties allow different areas of the string to interact with each other, causing it to fold up into a complex three-dimensional structure. That structure is essential for the protein’s function.




















+ There are no comments
Add yours